As the world has been learning about the precipitous and dangerous decline in bee populations, scientists are scrambling to probe deeper into the “why.” It’s generally recognized that habitat loss and degradation, increased use of agrochemicals, invasive pathogens, competition from alien species and poor management practices are each contributing factors. However, what’s not known is the extent and effect of each of these on various species of bees, and, further, the role that the interaction of the species in shared habitats and flower resources plays. It is supposed that each species will be affected by different degrees, because of differences in bee social organization, foraging and  nesting behavior, genetic diversity and other traits, but the hard science is not there. To take a closer look, Dr. Haw Chuan Lim and his Mason team of graduate and undergraduate students armed with a 4-VA2Mason grant are conducting groundbreaking research via their “High Throughput Bee Pathogen Survey.”
nesting behavior, genetic diversity and other traits, but the hard science is not there. To take a closer look, Dr. Haw Chuan Lim and his Mason team of graduate and undergraduate students armed with a 4-VA2Mason grant are conducting groundbreaking research via their “High Throughput Bee Pathogen Survey.”
In what may be the only study of its kind, the team is in the unique position not only to access,  but to develop state-of-the-art research techniques as they closely investigate extracted RNA and DNA from three bee species in Northern Virginia. Together, they are harnessing the bioinformatics and genomics capabilities at the Mason Sci-Tech campus while developing their own sequence capture probe-set to enable a comprehensive survey of pathogens and micro-parasites. They collaborate closely with Mason’s Rebecca Forkner and UVA’s T’ai Roulston. Both Forkner and Roulston have many years of experience in pollinator biology, using the Virginia Working Landscapes (VWL) program — the sites of the bee collection — and UVA’s Blandy Experimental Farm.
but to develop state-of-the-art research techniques as they closely investigate extracted RNA and DNA from three bee species in Northern Virginia. Together, they are harnessing the bioinformatics and genomics capabilities at the Mason Sci-Tech campus while developing their own sequence capture probe-set to enable a comprehensive survey of pathogens and micro-parasites. They collaborate closely with Mason’s Rebecca Forkner and UVA’s T’ai Roulston. Both Forkner and Roulston have many years of experience in pollinator biology, using the Virginia Working Landscapes (VWL) program — the sites of the bee collection — and UVA’s Blandy Experimental Farm.
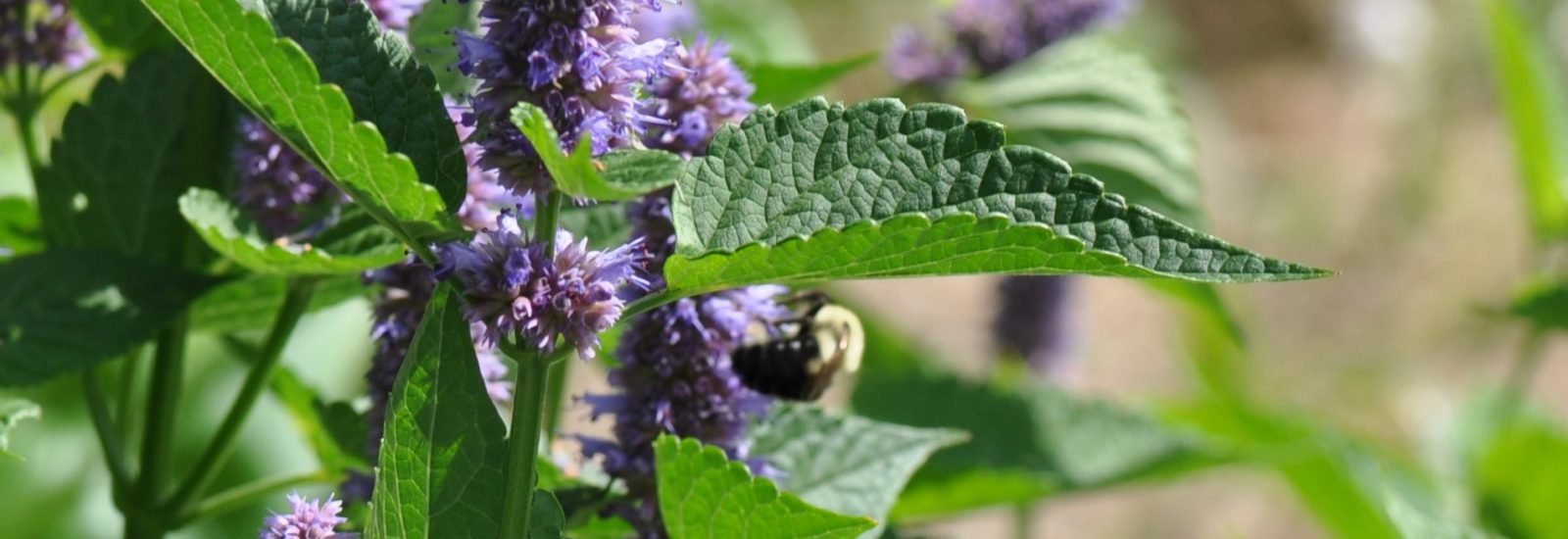
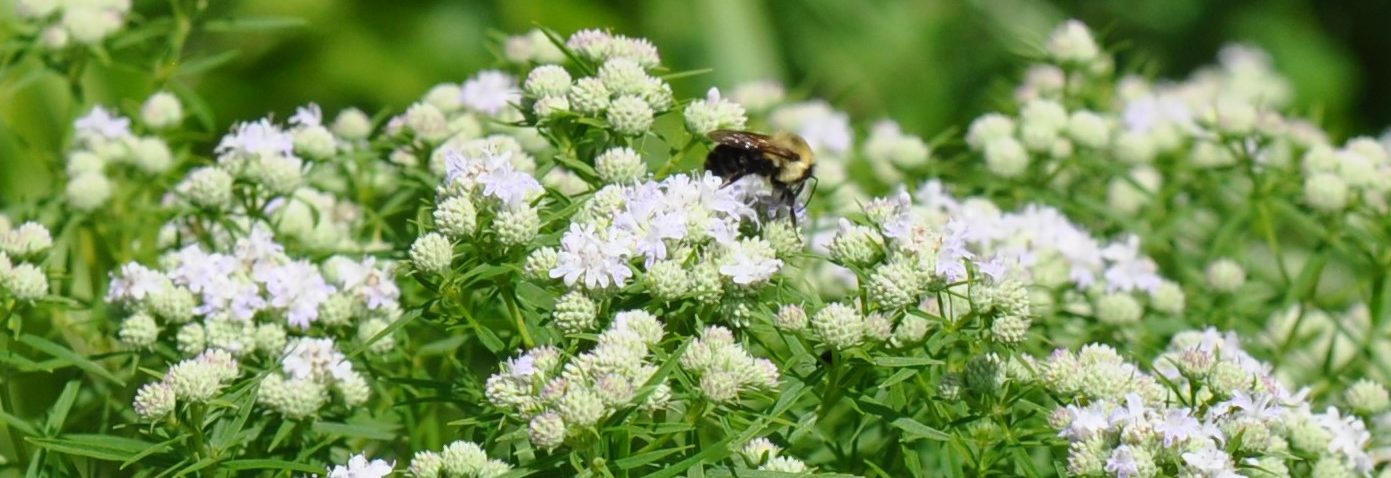
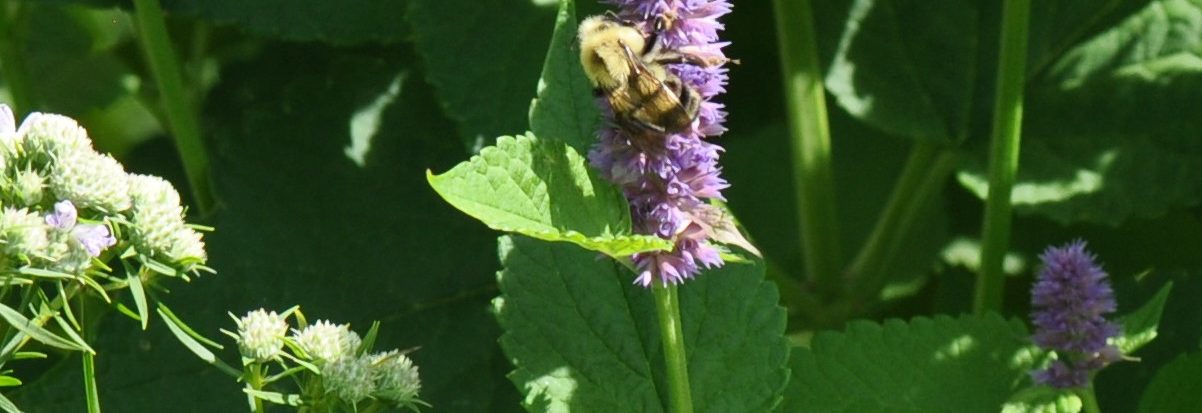
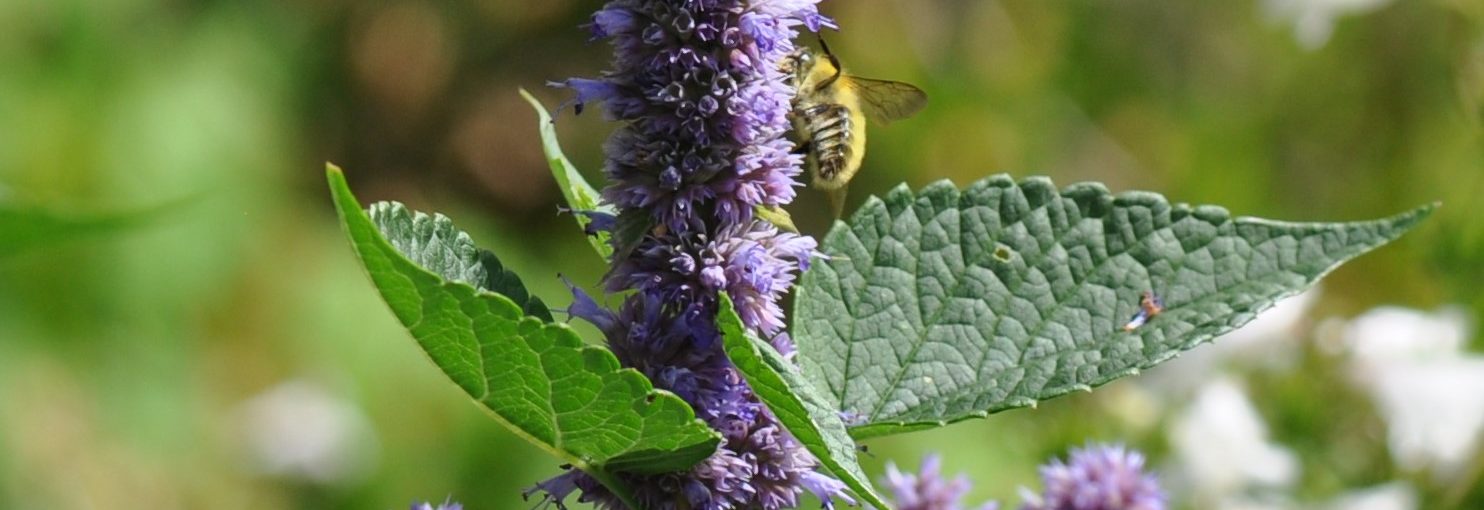
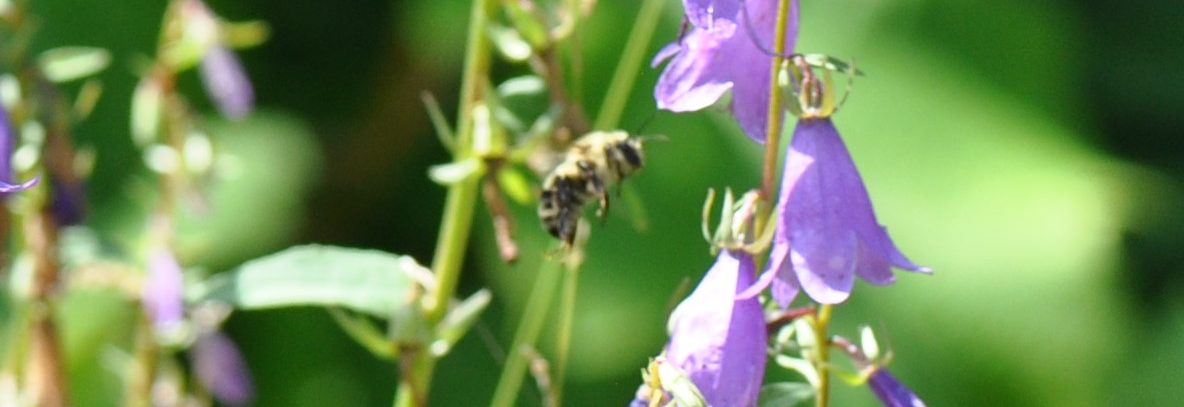
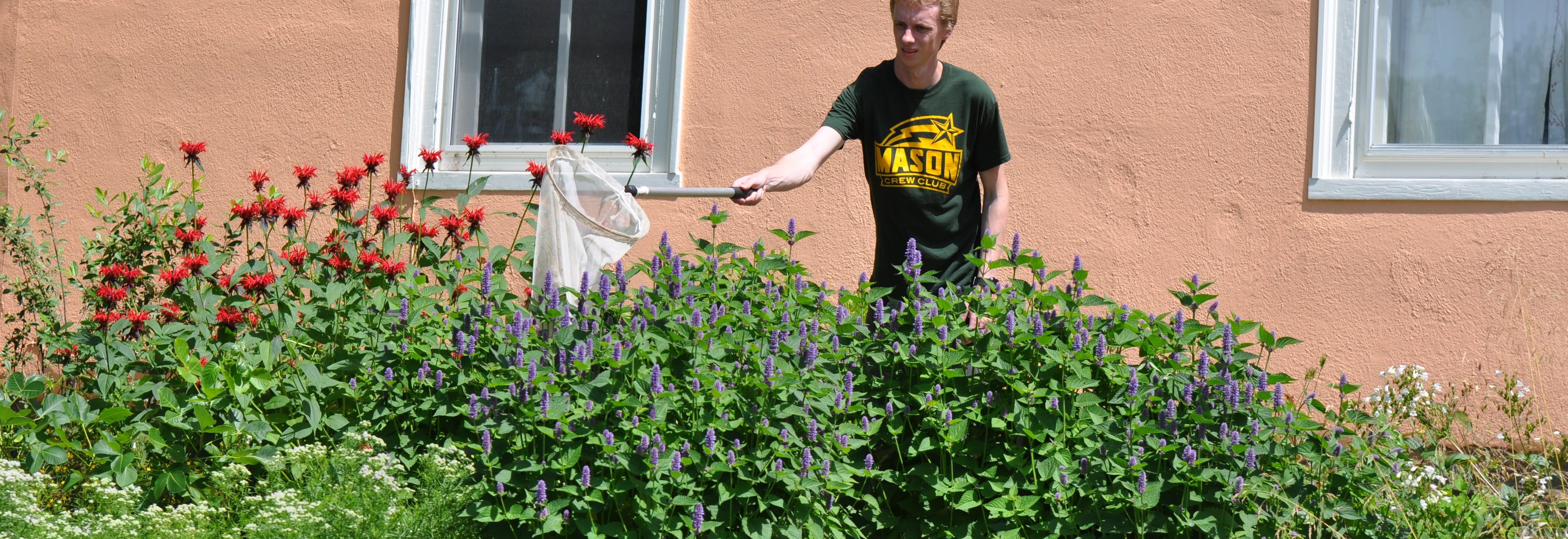
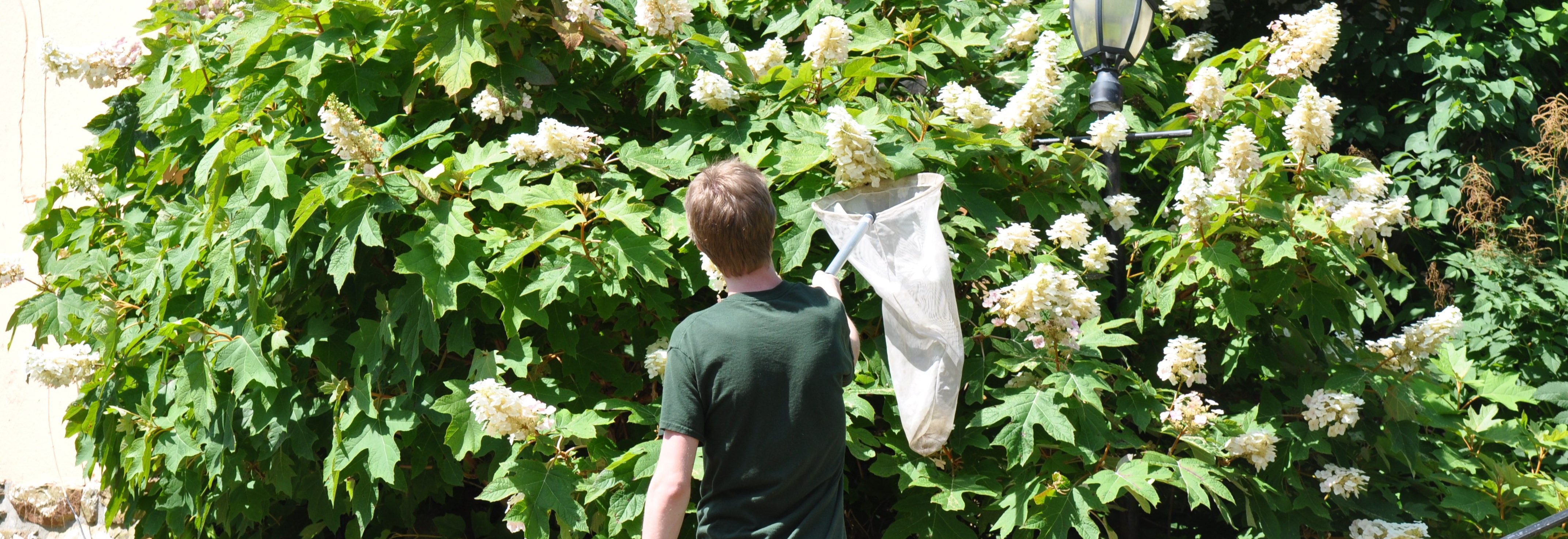
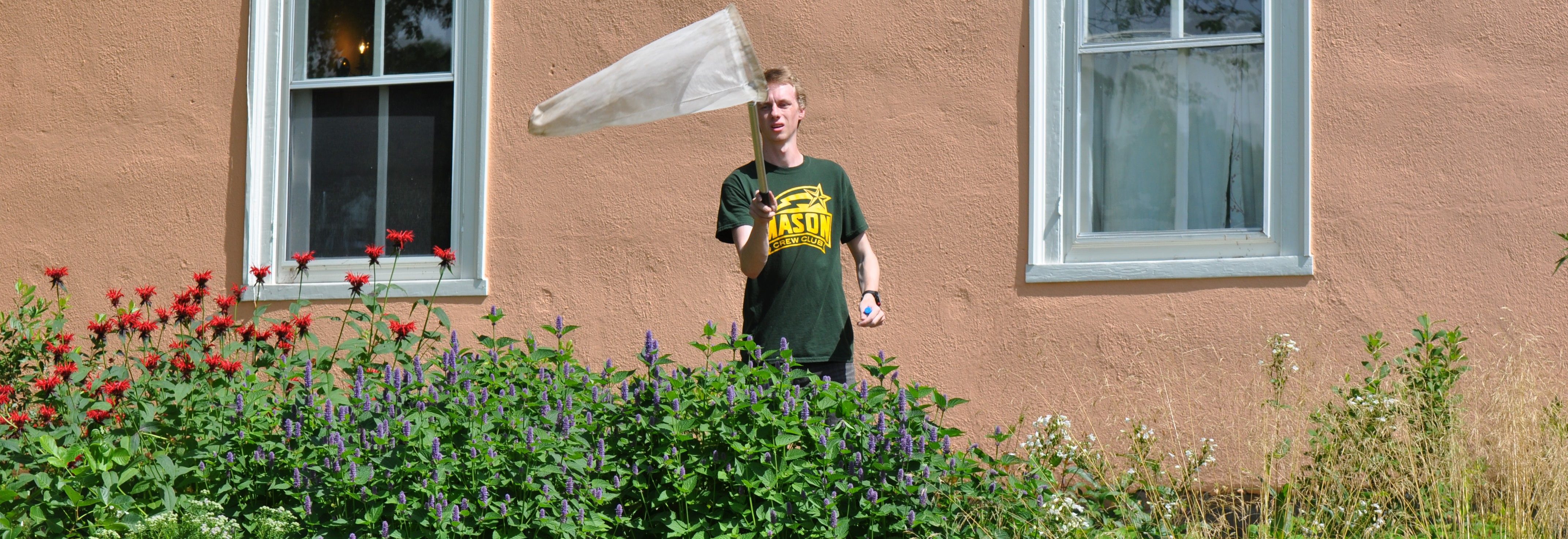
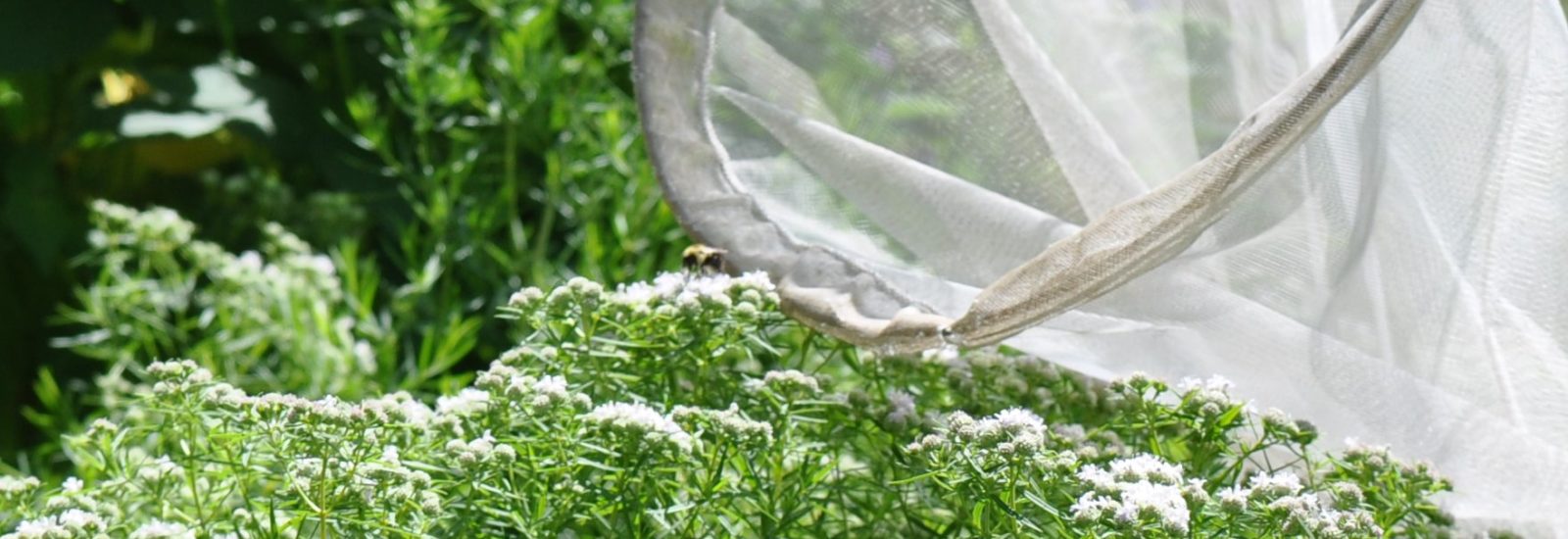
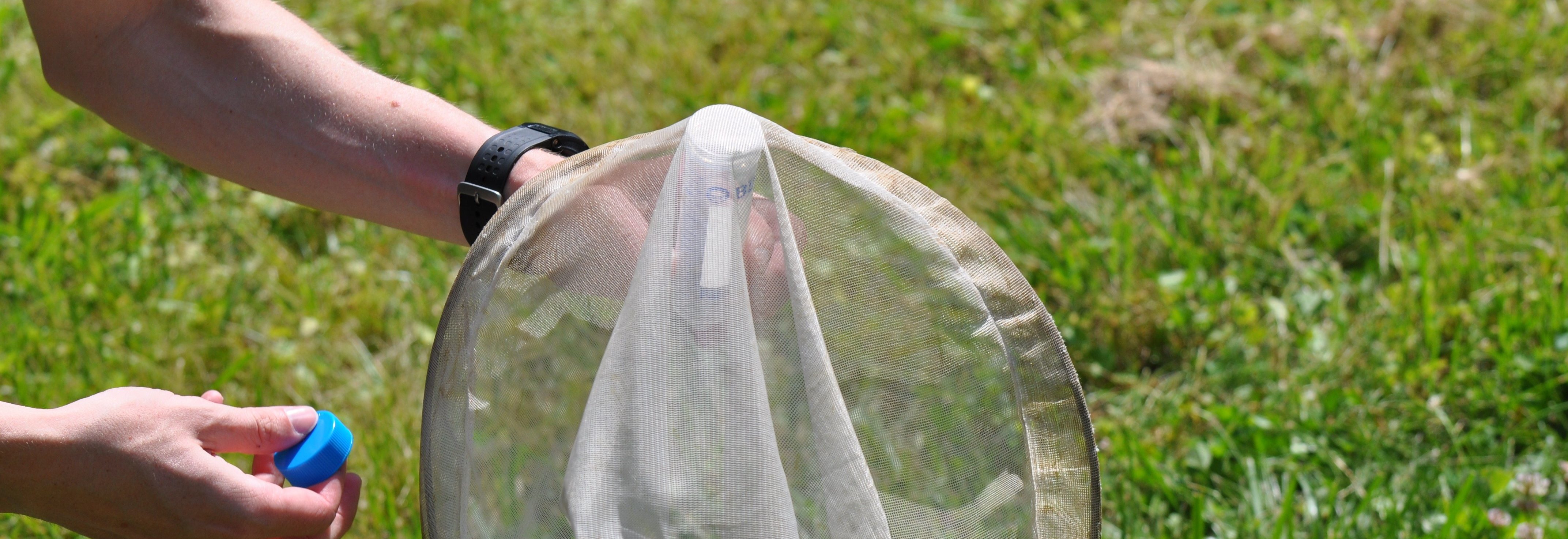
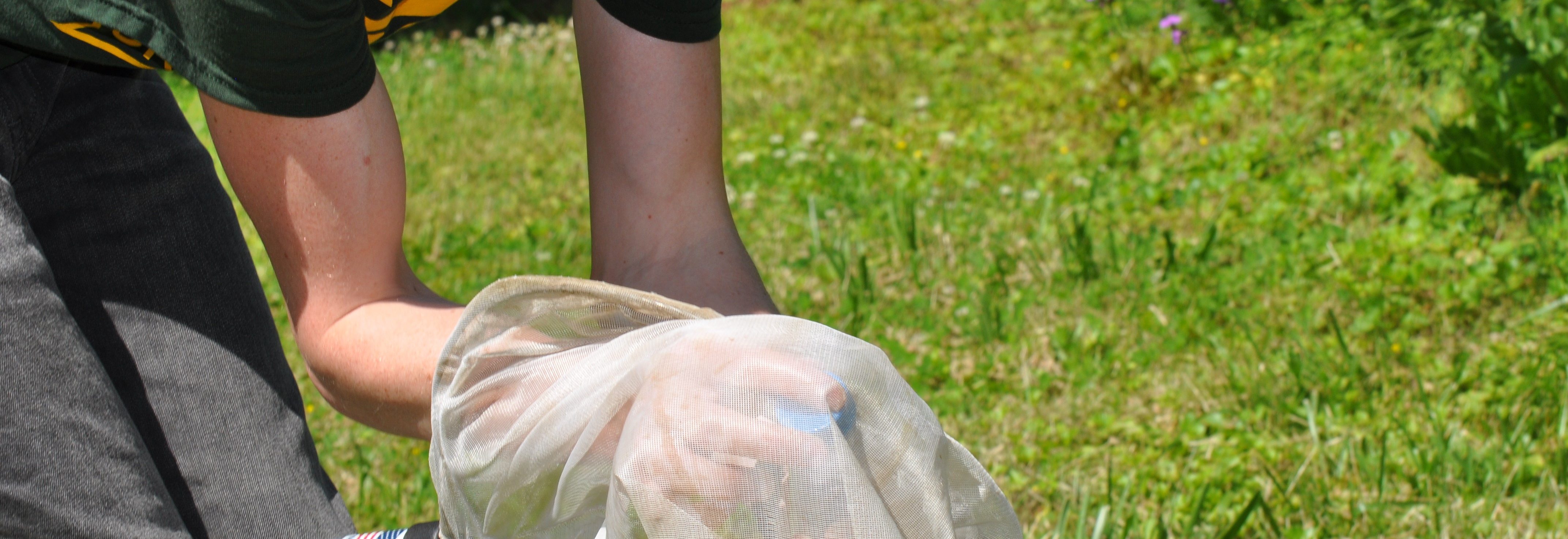
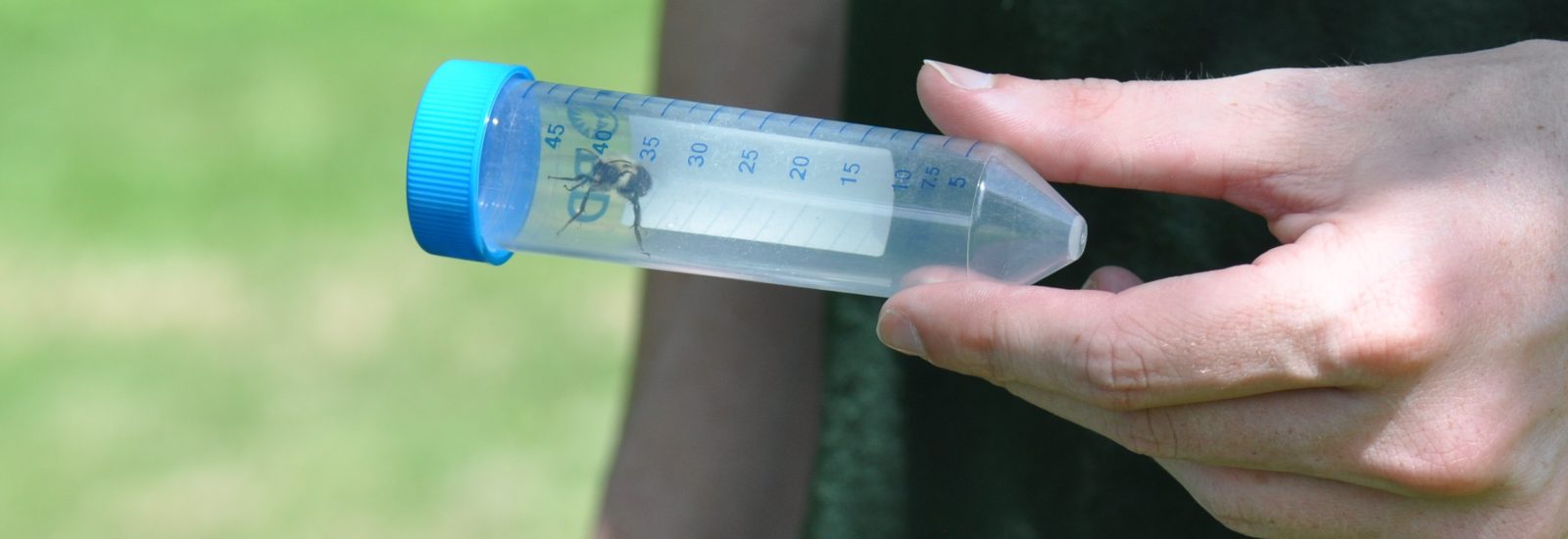
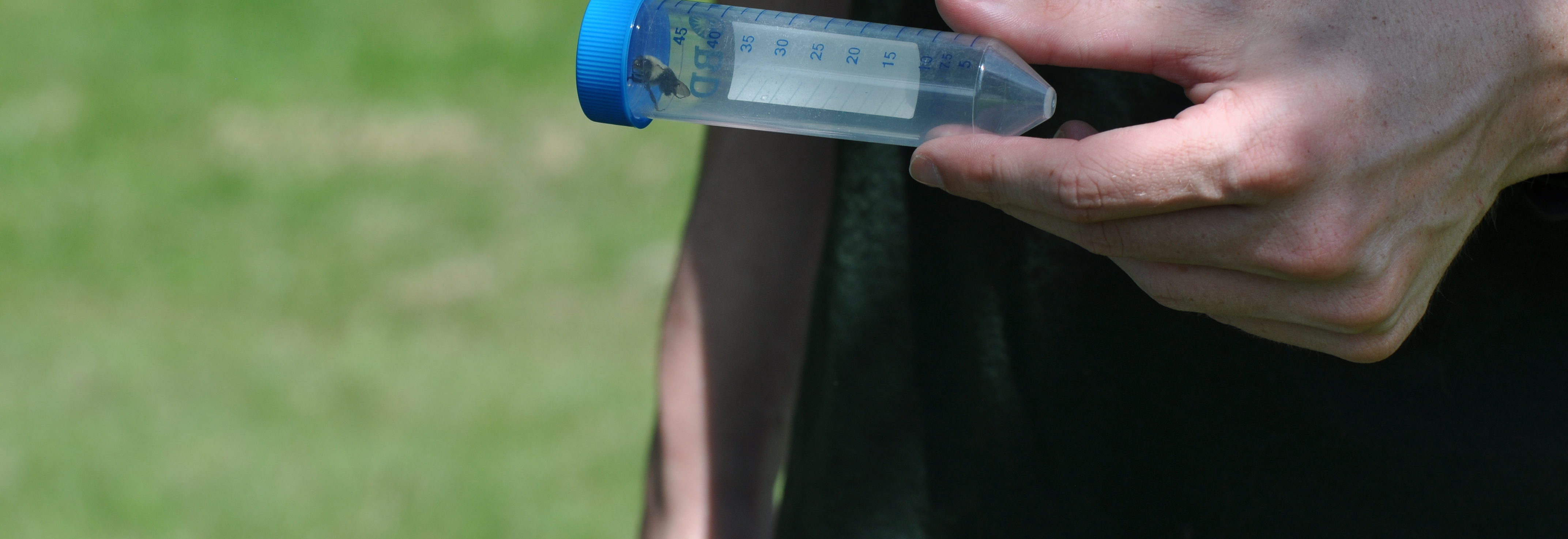
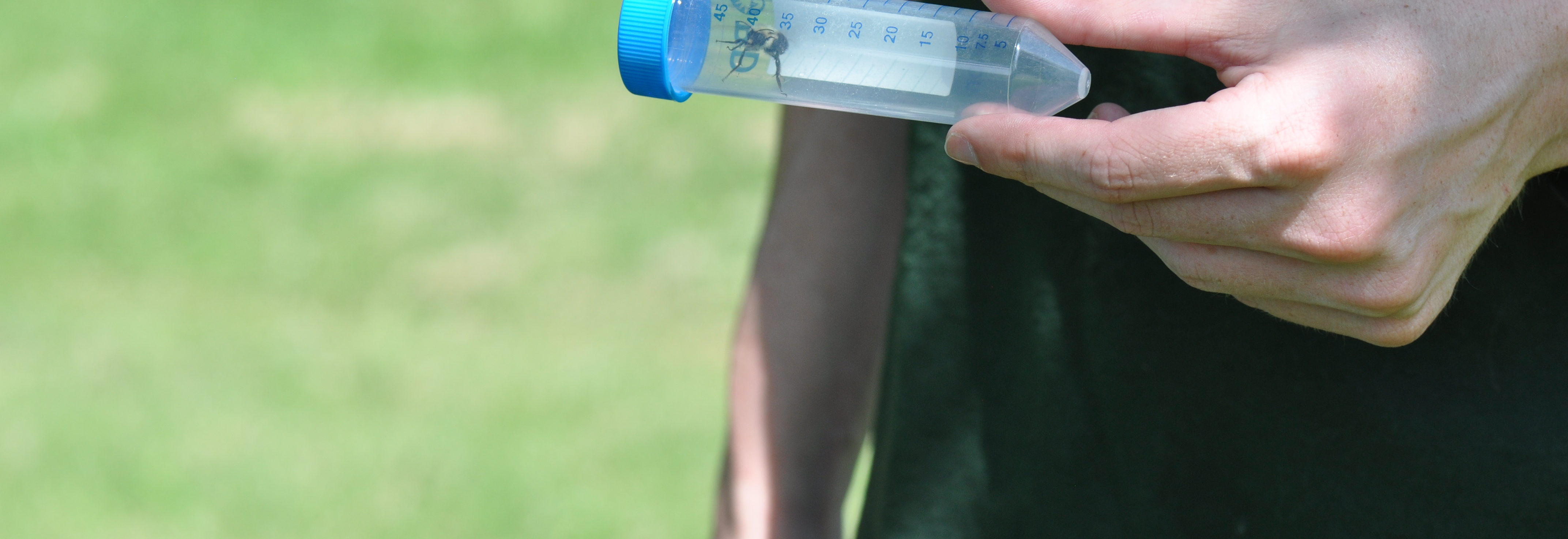
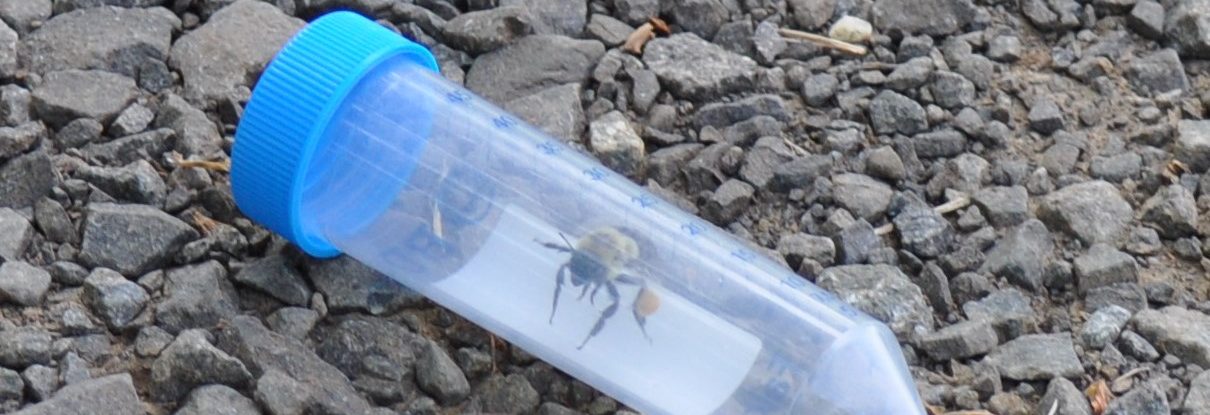
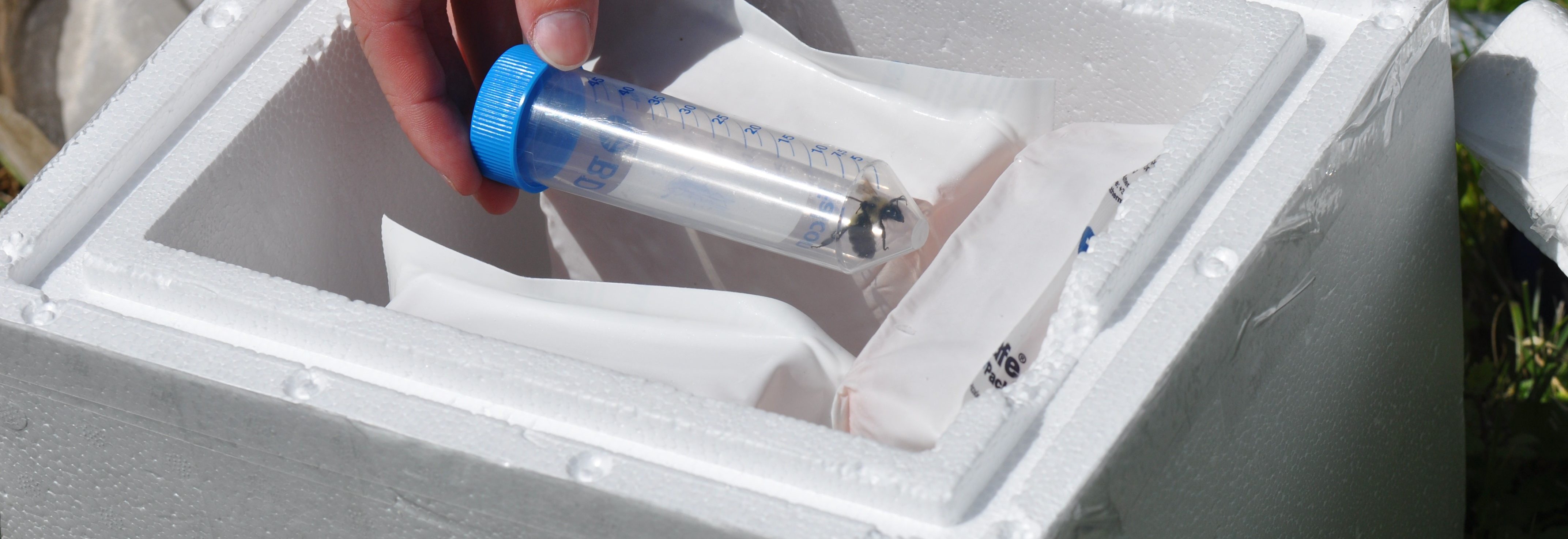
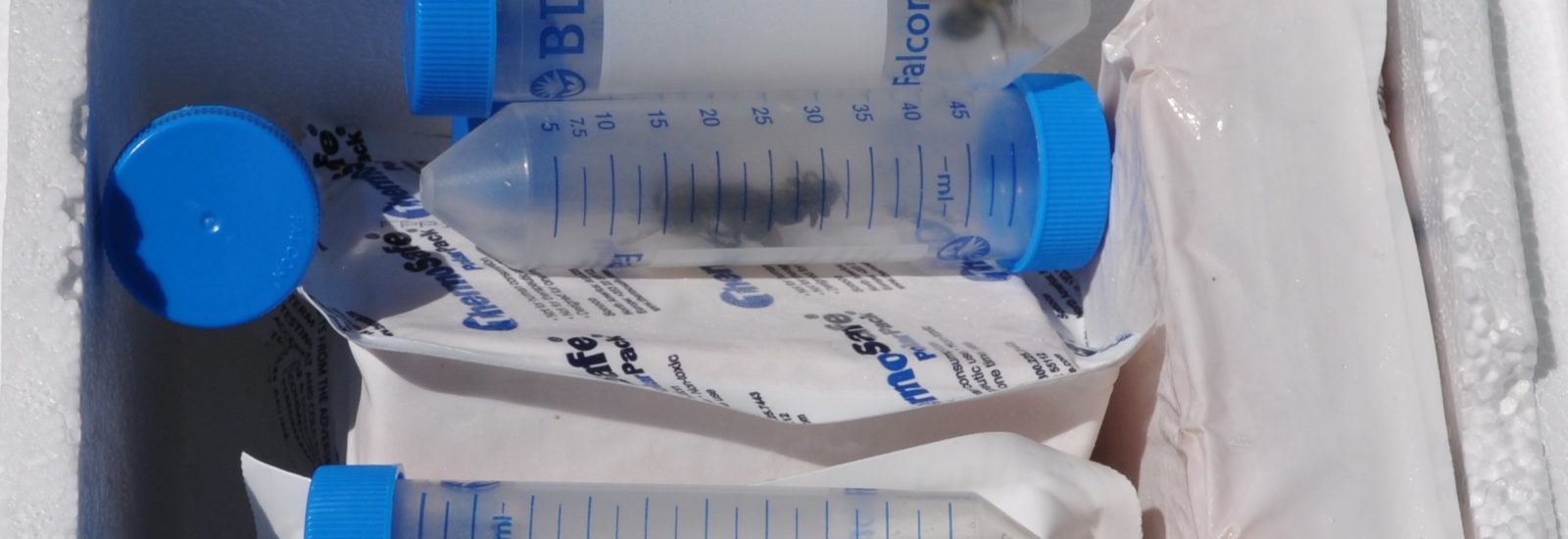
The team is specifically studying three groups of bees found in Northern Virginia – the European honeybee, the bumble bee and the mason bee – (no relation to George Mason University :+) To do this, they are collecting bees from nine VWL sites in the region, freezing and preserving each specimen using liquid nitrogen, and bringing them to the lab on the Sci-Tech campus, where they store them in a -80C freezer.
 When the initial series of specimens was harvested, master’s student David Lambrecht went
When the initial series of specimens was harvested, master’s student David Lambrecht went  into overdrive. “I’ve spent many long days and nights in this space,” notes Lambrecht as he removes a bee sample from the freezer. After freezing each specimen, the students on the project learned how to extract total RNA and total DNA from each specimen. By using techniques such as target sequence capture and polymerase chain reactions, they can then enrich for and sequence a variety of bee pathogens whose genomes are made up of RNA and DNA molecules.
into overdrive. “I’ve spent many long days and nights in this space,” notes Lambrecht as he removes a bee sample from the freezer. After freezing each specimen, the students on the project learned how to extract total RNA and total DNA from each specimen. By using techniques such as target sequence capture and polymerase chain reactions, they can then enrich for and sequence a variety of bee pathogens whose genomes are made up of RNA and DNA molecules.
Although the lab work is taxing, Lim notes that they’re making progress, and big progress at that. “This is the first time ever that a large-scale target enrichment and sequencing of RNA viruses have been implemented for bees in this region. More specifically, the prevalence of viruses is generally unknown for bees in this region.” explains Lim. “We have had to optimize lab protocols and bioinformatics analytical approaches.”
Collecting the baseline values and knowing the diversity and strain variation of pathogens can provide valuable information for the future of the bees, including:
- Being able to identify the pathogen responsible if bees in the region show signs for a particular disease. Conversely, it may be found that high prevalence or abundance of certain pathogen will not affect the bees, suggesting that they have developed resistance to the pathogen.
- Allowing scientists to target pathogens of interest and to conduct in vivo studies of the mechanisms of infection, as well as the
 immune responses of bees.
immune responses of bees. - Knowing whether managed bees (honey bees) are transmitting diseases to native bees will inform management practices, e.g. – keeping apiaries further away from native vegetation.
But while the initial lab work is buzzing away, the field work was thrown for a loop by Mother Nature. The total bee collections were hampered by the record setting rains in Northern Virginia this past year.
However, the persistent rains have not dampened the spirits of Lim and his team. They project that their research findings will not only shed light on critical information to help scientists better understand the bee populations and how to manage disease, stress and habitats; Lim also sees many valuable offshoots of this project for use in various upper division biology courses at Mason, and perhaps as a part of the Bioinformatics Concentration. Adds Lim, “My goal here is to help push along our bioinformatics and genomics program.”
And with the study still underway, Lim is already looking to the conclusion and beyond. Explains Lim, “Our results will be very relevant to the basic understanding of pollinator ecology, and management and conservation of bee populations. I foresee future funding from federal grant resources and private conservation organizations. Some of this lab work hasn’t been done before and it’s already opened up more research opportunities.”
——————————————————————————————————————————————————————-
A CLOSER LOOK…
… How the bees got to the lab. A story in pictures: