The science would not be easy. There were going to be long days and nights in the lab, countless tests to run and techniques to be tweaked. But the end game was intriguing; identify opportunities to affect modifications in the Francisella proteome, a bacterium categorized as a class “A” bioterrorism agent. Unlike its cousin, the more well-researched E. coli bacteria, many aspects of the Francisella proteome are not fully understood. Due to its potentially virulent nature, it is important to research and document the biochemistry of this organism in order to develop new therapies or vaccines.
While the van Hoek lab at the Mason Sci-Tech campus has been studying Francisella since 2005, they were interested to learn more about post-translational modifications (PTMs) – changes undergone by the protein when certain chemical groups are added. Specifically, Dr. Monique van Hoek, a professor in the School of Systems Biology, with a joint appointment to the National Center for Biodefense and Infectious Diseases at Mason, was interested in how the Francisella bacterium changes the activity of its proteins with the addition of the acetyl chemical group. Although van Hoek had years of experience in Francisella research, for this particular project she recognized that while her lab could make the protein/peptide samples, it was necessary to run a thorough molecular analysis of the samples to measure acetylation. That, she saw, could be done through a collaboration with another 4-VA partner university – Virginia Commonwealth, in their Chemical and Proteomics Mass Spectrometry Core Facility. It was that connection which paved the way for yet another groundbreaking 4-VA research project – “Critical post-translational modifications of the Francisella proteome.”

As van Hoek explains, not only did the research produce results, the grant also had a positive effect on students in her lab and faculty at VCU. “Real lives were changed — two great students graduated and went on to get good jobs,” notes van Hoek. The first student, Ekaterina (Kate) Marakosova, was a Ph.D. student at Mason who began the project with van Hoek working with the more virulent forms of Francisella. “Kate started on this project with me and developed techniques to identify protein acetylation. Kate has since gotten her doctorate and gone on to get a great job at the Food and Drug Administration,” says van Hoek. “Alex Ii is now working as a laboratory technician with me,” adds van Hoek. “In May, Alex defended her Master’s on another aspect of this project ‘Acetylation as a regulatory mechanism of chitinase activity in Francisella tularensis subsp. novidica.’”
Ii came to the van Hoek lab after starting her degree at VCU in Bioinformatics. “Once I got into this lab, I realized I really liked the work,” says Ii. “When I first started here, I was working with Kate and immediately jumped into the project. We were coming in at 5:00 am and often didn’t leave until 8:00 pm. The sample preparation was difficult, and we had to do a lot of troubleshooting, but it was worth it!” Ii was not only all in for the lab work, but with anything else that needed to be done, even driving samples to the lab in Richmond late at night.
van Hoek also notes that her collaborator at VCU, Dr. Kristina Nelson, points to the project in furthering her own research. Nelson received a 4-VA Complementary Grant for her part in the project. “With the complementary funding, we were able to purchase standards and columns in order to ensure that the instrument was operating at peak performance, to give the best data possible,” explains Nelson. “It was fascinating to be able to visualize the changes in the protein acetylation profile.”
“The new collaboration with Kristina was certainly another positive outcome of the 4-VA grant,” says van Hoek.
In addition to furthering the education and professional tracks of those on the project, the research was fruitful. The team has identified multiple Francisella proteins that are acetylated and look to be important in Francisella’s ability to infect hosts. To share the research, a poster was presented at the American Society for Microbiology meeting on biofilms and the manuscript has been submitted for potential publication.
While van Hoek notes there is still much more to be investigated with regard to the Francisella bacterium, which causes human disease in the US and in Europe, she credits the 4-VA@Mason grant for delivering these important results, and making such positive effects on the people and the science. van Hoek continues to study important questions of Francisella biology, such as which proteins are secreted by this bacterium and how they are exported. In fact, van Hoek and Nelson are now at work on another 4-VA collaborative research project on this very subject “Secreted Proteins of Francisella – a new understanding.” Stay tuned!



 the classroom to make it interesting and challenging,” explains Kim.
the classroom to make it interesting and challenging,” explains Kim. recognizes there is much to be understood about creating constructive introductions in the school setting. However, she is also keenly aware of a key flaw in the data used in the benchmark research – it is predominantly limited to students who are economically advantaged.
recognizes there is much to be understood about creating constructive introductions in the school setting. However, she is also keenly aware of a key flaw in the data used in the benchmark research – it is predominantly limited to students who are economically advantaged. Several months later, with that grant in hand, Garner identified an undergrad student, Tamera Toney, who was interested in the project and would be able to handle some of the data entry and management responsibilities. Toney worked on the project during her senior year at Mason. Garner saw that the 4-VA funding could provide a personal and professional pathway for Toney to enhance her studies. Toney recently graduated and will return in the fall to begin her master’s work in Psychology.
Several months later, with that grant in hand, Garner identified an undergrad student, Tamera Toney, who was interested in the project and would be able to handle some of the data entry and management responsibilities. Toney worked on the project during her senior year at Mason. Garner saw that the 4-VA funding could provide a personal and professional pathway for Toney to enhance her studies. Toney recently graduated and will return in the fall to begin her master’s work in Psychology.

 rehearsed and studied the music within its historical context. And similar to good musical composition, RtA worked to a crescendo. For the RtA team, that was a spring evening in Charlottesville when the team of researchers, performers (musicians and singers), videographers and archivists, librarians, faculty and more joined together in UVA’s Colonnade Club Garden Room to fully embrace 17 pieces of WWI music. From “K-K-K Katy” to “Over There” to “Oh How I Hate to Get Up in the Morning” the Colonnade Room sprang to life — circa 1918.
rehearsed and studied the music within its historical context. And similar to good musical composition, RtA worked to a crescendo. For the RtA team, that was a spring evening in Charlottesville when the team of researchers, performers (musicians and singers), videographers and archivists, librarians, faculty and more joined together in UVA’s Colonnade Club Garden Room to fully embrace 17 pieces of WWI music. From “K-K-K Katy” to “Over There” to “Oh How I Hate to Get Up in the Morning” the Colonnade Room sprang to life — circa 1918.


 Team member Psyche Z. Ready, assisted by Joyce P. Johnston, took the lead in adapting Mason Journals’ iteration of the Open Journal System (OJS) to meet the needs of English 302 OER collection authors, reviewers, and users. Each item in the new, public-facing collection includes an abstract, instructor’s notes, and creative-commons licensed curricular materials – assignments, activities, and/or background readings – created, adapted, or curated for use in English 302. The OJS platform eases the review process, and also allows user-friendly features such as keyword searching.
Team member Psyche Z. Ready, assisted by Joyce P. Johnston, took the lead in adapting Mason Journals’ iteration of the Open Journal System (OJS) to meet the needs of English 302 OER collection authors, reviewers, and users. Each item in the new, public-facing collection includes an abstract, instructor’s notes, and creative-commons licensed curricular materials – assignments, activities, and/or background readings – created, adapted, or curated for use in English 302. The OJS platform eases the review process, and also allows user-friendly features such as keyword searching. ead PI Moissa Fayissa, PhD. conjectured that this might just be the path for the team to pursue: He believed their current text and lab books were subpar and incomplete as a match for their course. Fayissa saw the need to provide only top-notch materials for this intensive class — which is offered in three sessions in the fall semester and two sessions in the spring semester. Additionally, Fayissa worried about the cost of their then-current textbook. At more than $250, this was a high price to ask students to pay.
ead PI Moissa Fayissa, PhD. conjectured that this might just be the path for the team to pursue: He believed their current text and lab books were subpar and incomplete as a match for their course. Fayissa saw the need to provide only top-notch materials for this intensive class — which is offered in three sessions in the fall semester and two sessions in the spring semester. Additionally, Fayissa worried about the cost of their then-current textbook. At more than $250, this was a high price to ask students to pay. , “The materials search included looking at printed laboratory manuals and online open resources. When we could not find enough information online for the experiment, we referred to the previous laboratory manual and cited the lab manual as the reference. The instructions and background materials found online were rewritten to suit our needs.”
, “The materials search included looking at printed laboratory manuals and online open resources. When we could not find enough information online for the experiment, we referred to the previous laboratory manual and cited the lab manual as the reference. The instructions and background materials found online were rewritten to suit our needs.”
 telepresence technology at each of the 4-VA schools.
telepresence technology at each of the 4-VA schools. learning and improvement” was an instant success, with 168 conference registrants representing 50 organizations: 31 universities, 15 community colleges, and 4 professional organizations. The event was funded by a 4- VA Collaborative Research Grant and organized by the nonprofit Virginia Assessment Group.
learning and improvement” was an instant success, with 168 conference registrants representing 50 organizations: 31 universities, 15 community colleges, and 4 professional organizations. The event was funded by a 4- VA Collaborative Research Grant and organized by the nonprofit Virginia Assessment Group. description of the gold standard, randomized control trial, followed by a “let’s get real” section highlighting the real-world data collection challenges that assessment practitioners face. Participants grappled with how to make appropriate inferences from the data collection designs that are possible given common constraints. The morning concluded with participants from each location providing suggestions for ways of dealing with practical challenges related to data collection.
description of the gold standard, randomized control trial, followed by a “let’s get real” section highlighting the real-world data collection challenges that assessment practitioners face. Participants grappled with how to make appropriate inferences from the data collection designs that are possible given common constraints. The morning concluded with participants from each location providing suggestions for ways of dealing with practical challenges related to data collection. and Gianina Baker (National Institute for Learning Outcomes Assessment – NILOA). Participants viewed a video produced by Jillian Kinzie (NILOA), illustrating examples and rationale for presenting assessment findings that tell the story of student learning. Participants engaged in an activity in which they tailored a data report to a particular stakeholder audience. Gianina Baker closed the afternoon, providing reflections and suggestions for effective evidence-based reporting.
and Gianina Baker (National Institute for Learning Outcomes Assessment – NILOA). Participants viewed a video produced by Jillian Kinzie (NILOA), illustrating examples and rationale for presenting assessment findings that tell the story of student learning. Participants engaged in an activity in which they tailored a data report to a particular stakeholder audience. Gianina Baker closed the afternoon, providing reflections and suggestions for effective evidence-based reporting.
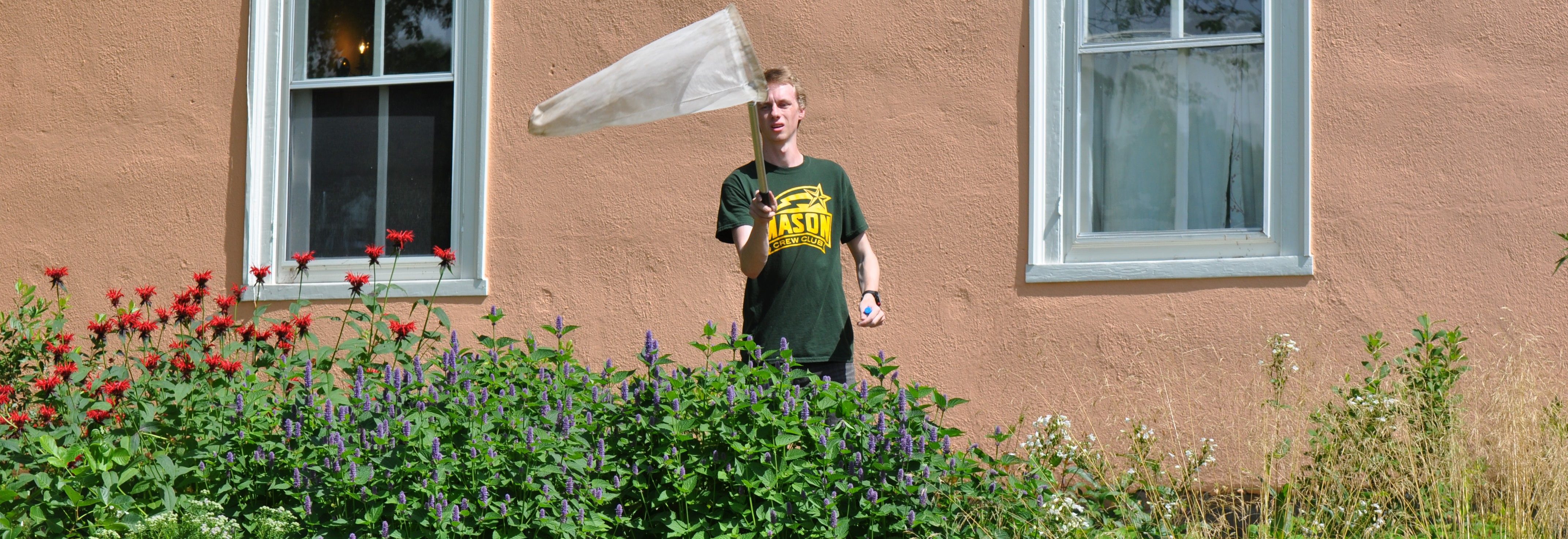
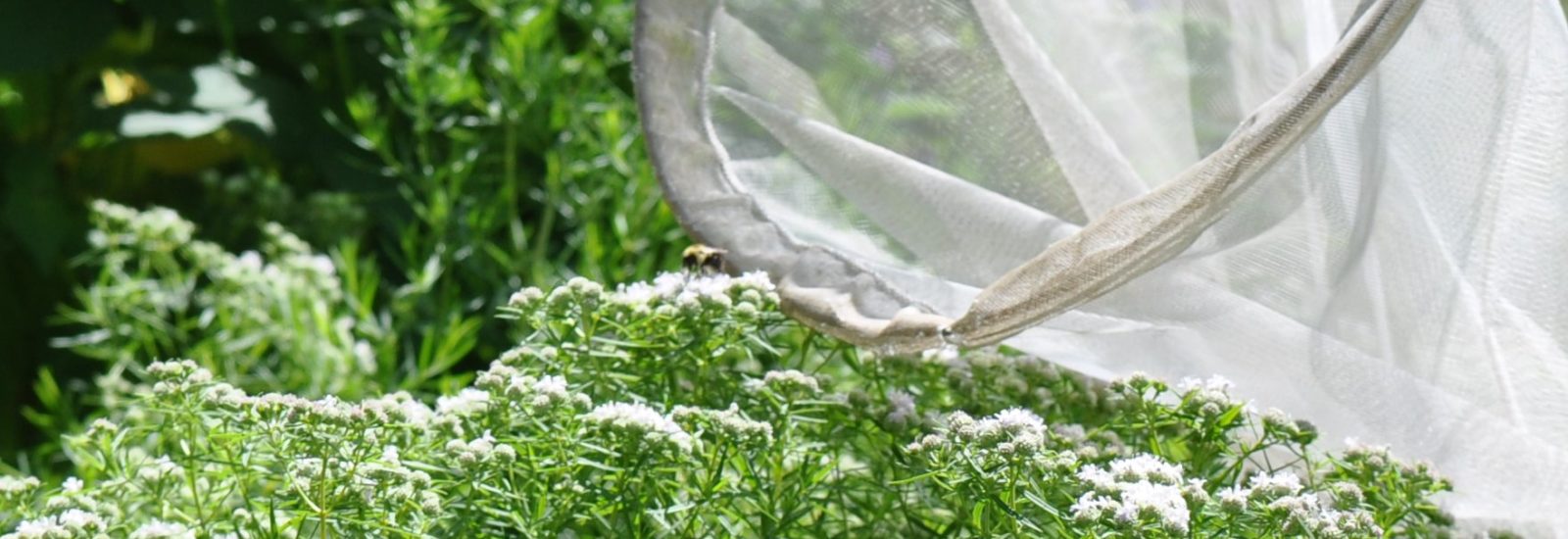
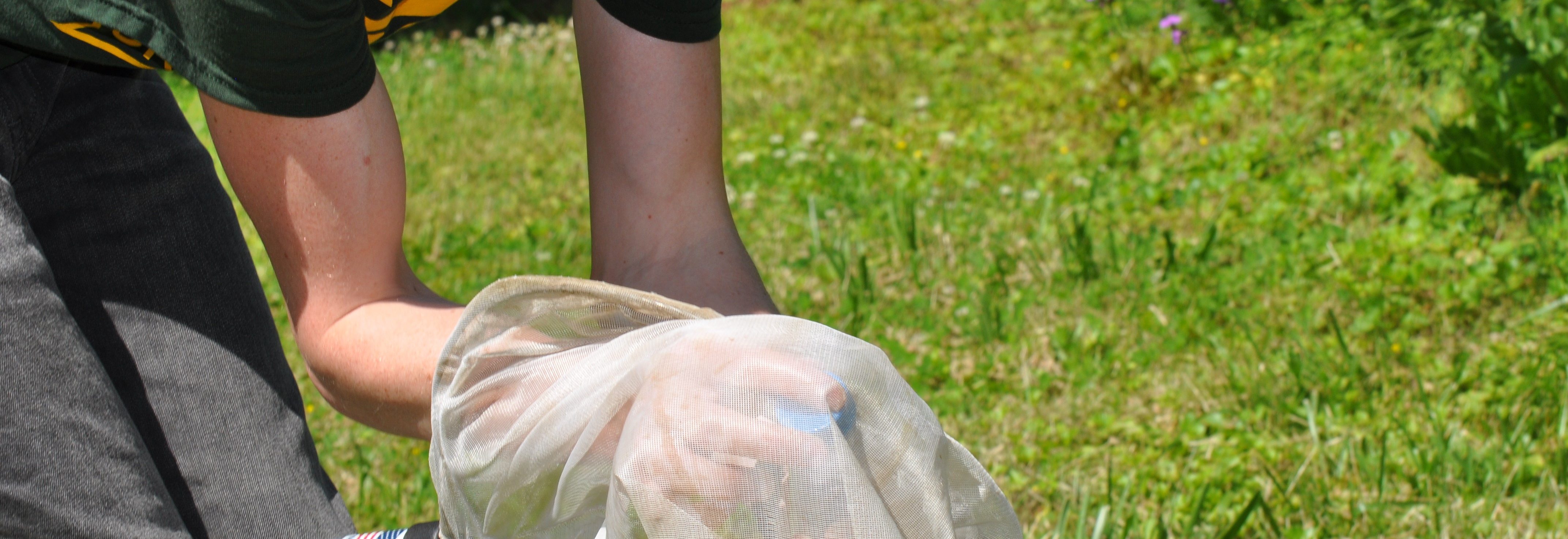
 nesting behavior, genetic diversity and other traits, but the hard science is not there. To take a closer look, Dr. Haw Chuan Lim and his Mason team of graduate and undergraduate students armed with a 4-VA2Mason grant are conducting groundbreaking research via their “High Throughput Bee Pathogen Survey.”
nesting behavior, genetic diversity and other traits, but the hard science is not there. To take a closer look, Dr. Haw Chuan Lim and his Mason team of graduate and undergraduate students armed with a 4-VA2Mason grant are conducting groundbreaking research via their “High Throughput Bee Pathogen Survey.” but to develop state-of-the-art research techniques as they closely investigate extracted RNA and DNA from three bee species in Northern Virginia. Together, they are harnessing the bioinformatics and genomics capabilities at the Mason Sci-Tech campus while developing their own sequence capture probe-set to enable a comprehensive survey of pathogens and micro-parasites. They collaborate closely with Mason’s Rebecca Forkner and UVA’s T’ai Roulston. Both Forkner and Roulston have many years of experience in pollinator biology, using the Virginia Working Landscapes (VWL) program — the sites of the bee collection — and UVA’s Blandy Experimental Farm.
but to develop state-of-the-art research techniques as they closely investigate extracted RNA and DNA from three bee species in Northern Virginia. Together, they are harnessing the bioinformatics and genomics capabilities at the Mason Sci-Tech campus while developing their own sequence capture probe-set to enable a comprehensive survey of pathogens and micro-parasites. They collaborate closely with Mason’s Rebecca Forkner and UVA’s T’ai Roulston. Both Forkner and Roulston have many years of experience in pollinator biology, using the Virginia Working Landscapes (VWL) program — the sites of the bee collection — and UVA’s Blandy Experimental Farm. When the initial series of specimens was harvested, master’s student David Lambrecht went
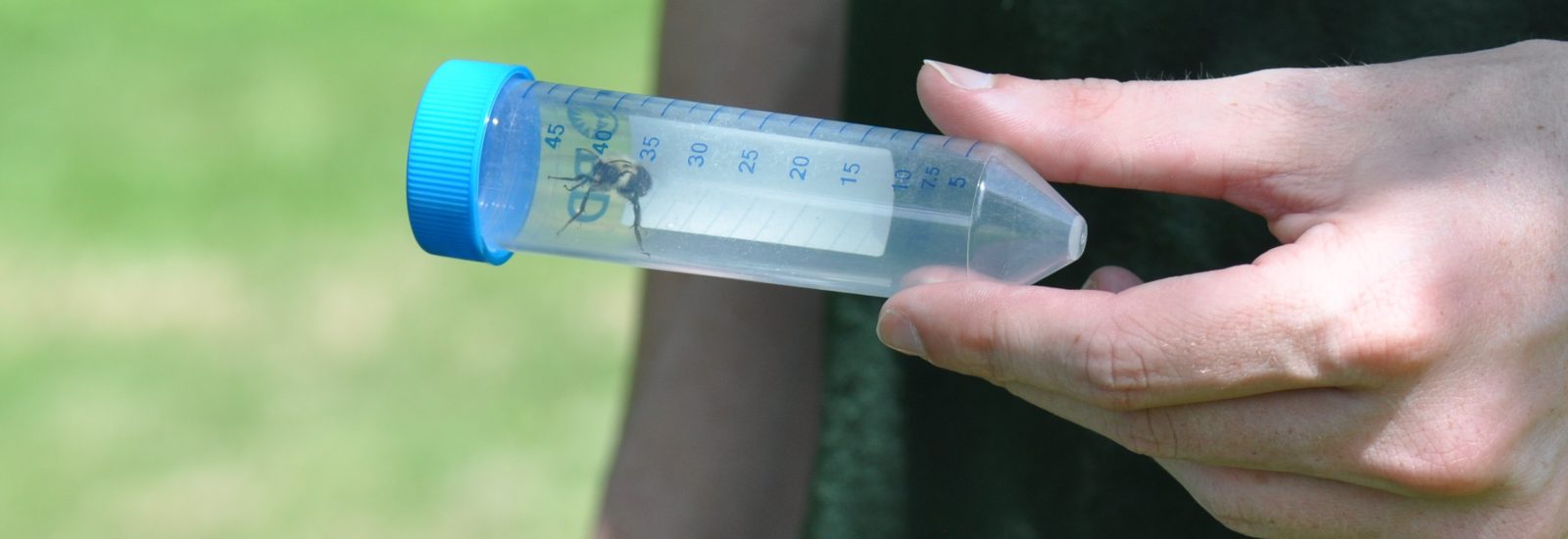
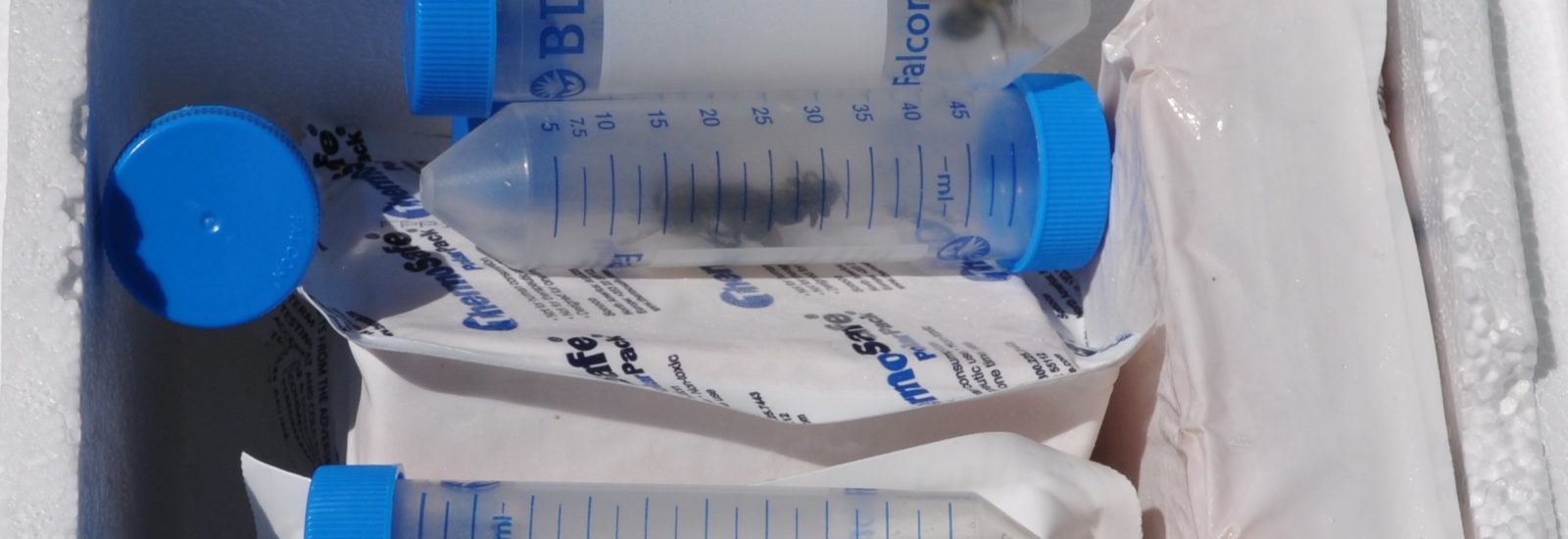
When the initial series of specimens was harvested, master’s student David Lambrecht went  into overdrive. “I’ve spent many long days and nights in this space,” notes Lambrecht as he removes a bee sample from the freezer. After freezing each specimen, the students on the project learned how to extract total RNA and total DNA from each specimen. By using techniques such as target sequence capture and polymerase chain reactions, they can then enrich for and sequence a variety of bee pathogens whose genomes are made up of RNA and DNA molecules.
into overdrive. “I’ve spent many long days and nights in this space,” notes Lambrecht as he removes a bee sample from the freezer. After freezing each specimen, the students on the project learned how to extract total RNA and total DNA from each specimen. By using techniques such as target sequence capture and polymerase chain reactions, they can then enrich for and sequence a variety of bee pathogens whose genomes are made up of RNA and DNA molecules.
 immune responses of bees.
immune responses of bees.